Illumina Microbial Amplicon Prep (IMAP)
New library prep solutions and a data analysis software for One Health genomic surveillance and research. The *Illumina Microbial Amplicon Prep* product line, launching today, is built on the foundation of the robust and popular COVIDSeq Assay workflow.
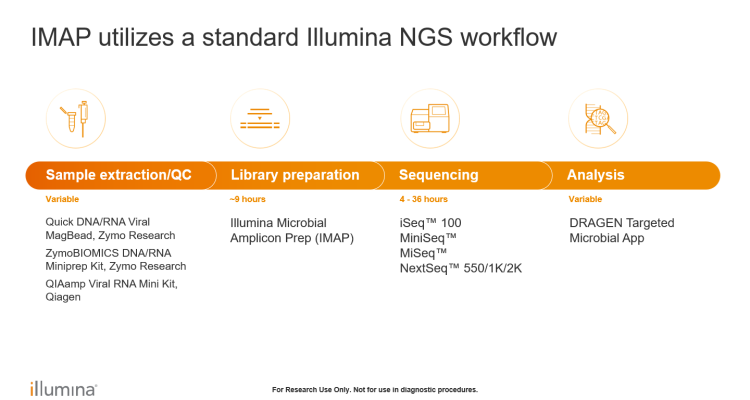
Illumina Microbial Amplicon Prep (iMAP) is an amplicon-based library preparation kit built on the same chemistry as COVIDSeq. Library preparation is fast and simple, with libraries compatible with nearly all Illumina sequencing systems. It can be used for various infectious disease, public health surveillance, and microbial research applications, including viral whole-genome sequencing, antimicrobial resistance marker analysis, bacterial and fungal identification, and more.
Expand Your Microbial Research Capabilities
Get sequencing results with exceptional data quality and accuracy at any scale with low hands-on time:
- Target variants with ready-to-use or lab-designed oligos
- Use a multiplexed, PCR-based workflow
- Interpret data with simple analysis solutions
- Sequence a broad range of samples
The kit can be used with custom, published, or commercially available primer sets (primer oligos are not included) and is compatible with RNA or DNA extracted from sources such as cultures, swabs (nasopharyngeal, skin, and nasal), wastewater, and more. Analysis can be performed using the DRAGEN Targeted Microbial App on Basespace Sequence Hub with pre-loaded targets. Files can also be easily uploaded for custom analyses.
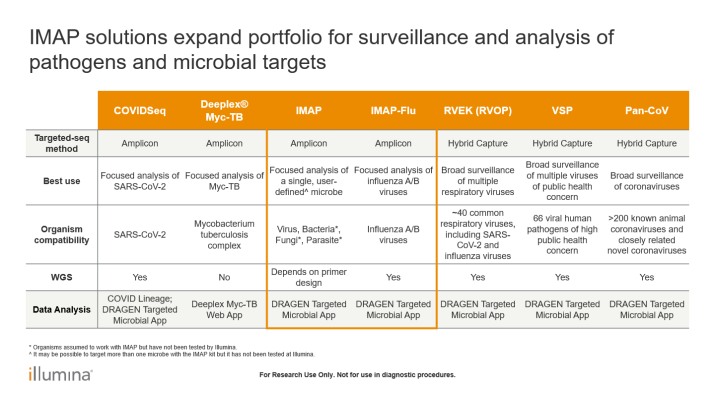
IMAP
- Versatile library preparation solution for public health surveillance and microbiology research
- Supports user-designed primers to sequence pathogens of public health concern
- Enables high-quality genome-wide coverage across multiple microbial species
- Accommodates DNA and RNA inputs from a range of sample sources and types
IMAP-flu
- Streamlined library preparation solution for influenza virus research and public health surveillance
- Enables rapid, unbiased, whole-genome coverage of influenza A and B virus strains
- Accommodates RNA inputs from a range of sample types and viral loads
- Simplifies whole-genome analysis and variant interpretation